In 2008, I was perfectly healthy. By the end of 2009, at the age of 25, I had been diagnosed with PCOS, PMDD, grade three anemia, cirrhosis, a kidney mass, reflux, peripheral neuropathy, optic neuritis, bacterial vaginosis, SIDS, chronic edema, endometriosis, and celiac disease. I gained 50 pounds in three months. I looked like a giant gray balloon.
Because of the sudden onset of all of these conditions, I asked myself what I thought was going to be a simple question: what if these are not separate conditions at all, but symptoms of something else, something bigger? And if so, symptoms of what?
Dysbiosis. Obviously.
That question, asked out of complete and utter desperation, led me to the microbiome literature in 2010. And what I found was that my conditions had overlapping signatures. When I pieced those signatures together and got a more coherent pattern, I began figuring out how to decrease the pathogens and support the more beneficial ones. I read everything I could get my hands on. At the time, there were only a dozen or so relevant papers. Somehow, that was enough.
What follows is not what I did. It is how I did what I did, and what that looks like when applied to a single condition: endometriosis. This is a complete walkthrough of the reasoning method I have been developing and refining for 15 years.
What a microbiome signature actually is (and is not)
Before walking through the method, it is necessary to address what most researchers mean when they say "microbiome signature" and why that definition is insufficient.
Microbiome signatures are commonly reduced to lists of differentially abundant disease-associated taxa, judged by reproducibility and statistical significance. Because of this definition, many researchers have argued that a definitive microbiome signature is completely unattainable given regional differences in microbiota and inconsistent methods across studies.
My contention is that a true microbiome signature has to be more nuanced than that. Imposing standardization to resolve variability risks systematic bias. A microbiome signature must ultimately provide a framework that incorporates and tolerates regional differences and inconsistent methods. Taxon-centric framing is rigid, non-intuitive, and makes translational research unnecessarily difficult.
The question I keep returning to is this: what is the point of this data if it is not translational? What is the point if it does not give us a first-principles understanding of a disease? What is the point if it does not help us connect the dots, form hypotheses, or develop novel interventions? And most importantly, what is the point if it cannot even support predictive modeling or ultimately save lives?
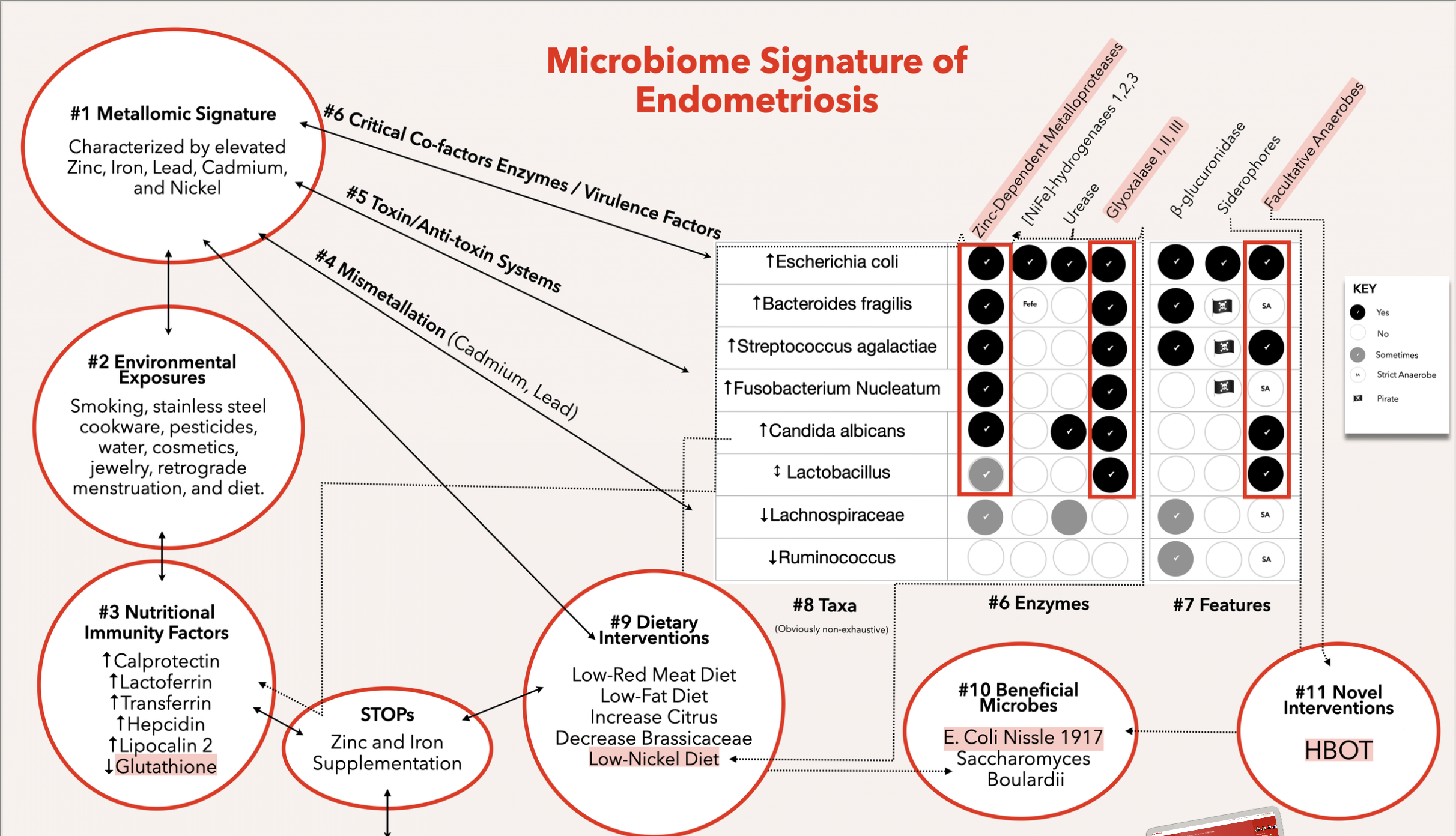
Step 1: The metallomic signature
The method begins not with taxa but with metals. The metallomic signature of endometriosis is characterized by elevated zinc, iron, cadmium, lead, and nickel in the tissues.
This is the starting point because the metal pattern constrains everything downstream. It determines which organisms can persist, which virulence factors will be expressed, and which interventions will or will not work. Metal-level analysis always precedes taxon-level analysis. For the broader research synthesis on metals, selection pressure, and pathogens, see microbial metallomics and heavy metal contamination.
Step 2: Environmental exposures
The environmental sources feeding the metallomic signature include smoking, stainless steel cookware, pesticides (especially organophosphates), water, cosmetics, jewelry, retrograde menstruation (which is technically a micro-environment exposure), and diet, which is the largest contributor of all.
Step 3: Nutritional immunity
Here is where the method diverges from conventional thinking. The host is not passively accumulating these metals. The host is actively responding to them through nutritional immunity, the deliberate restriction of metal availability as a defense mechanism.
In endometriosis, the following host defense markers are elevated: calprotectin (chelates and sequesters zinc), lactoferrin (chelates iron but also nickel), transferrin (chelates iron locally), hepcidin (increased in the presence of high iron, decreased during iron deficiency), and lipocalin-2 (a siderophore-binding protein). All elevated.
Glutathione, however, is decreased. This is a critical finding because among all the nutritional immunity factors listed, glutathione is the only one that can neutralize cadmium and lead.
This tells us something specific: the host is successfully defending against zinc and iron-seeking pathogens but is losing the fight against cadmium and lead toxicity. That asymmetry matters.
Step 4: Mismetallation
Cadmium and lead both enter cells through calcium channels, displacing correct cofactors like zinc or iron from enzymes. This leads to hypoxia. When cadmium and lead are combined in the environment, they produce a synergistic effect, generating even more oxidative stress than either metal alone.
Step 5: Pathogenic selection
Now the method arrives at taxa, but through the lens of the metallomic environment rather than through differential abundance alone.
Taxa with more robust efflux pumps or those that can survive in iron-rich, pro-inflammatory environments will outcompete taxa that cannot. The taxa that persist often use zinc, iron, and nickel as critical cofactors for metal-dependent enzymes.
The key enzymes are: zinc-dependent metalloproteinases (which help bacteria degrade host tissues and inhibit the host immune system), [NiFe]-hydrogenases (iron and nickel-dependent enzymes that decrease local oxygen saturation, creating a hypoxic environment suitable for strict or facultative anaerobes), nickel-dependent urease (which increases local pH via ammonia and CO2, disrupting the tightly regulated pH that vaginal lactobacilli need to survive), and nickel-dependent glyoxalase (which allows pathogens to persist against neutrophils in the blood).
Additionally, beta-glucuronidase positivity enables increased hepatic estrogen recirculation from estrogen glucuronide conjugation, a hallmark of endometriosis. Siderophore expression allows pathogens to chelate and uptake iron from their host or environment. And the presence of facultative anaerobes creates a feedback loop: in order to survive in a low-oxygen state, they switch to fermentative metabolism, but then the lesion microenvironment becomes acidic, zinc-rich, iron-rich, and pro-inflammatory, which is growth-promoting. This further amplifies lesion survival and invasion, perpetuating the condition.
Step 6: The taxa, mapped to the framework
With the metallomic environment and enzyme landscape established, the specific taxa now make sense.
E. coli is disproportionately high in endometriosis. It has all of the zinc, iron, and nickel-dependent enzymes, plus the mechanistic ability for estrogen deconjugation. It ticks every box in the framework above.
Bacteroides fragilis utilizes zinc as a critical cofactor for its main virulence factor (the fragilysin toxin, BFT). It expresses a super iron-dependent hydrogenase and even pirates iron from other microbes. It expresses beta-glucuronidase. It is a strict anaerobe, so its presence indicates hypoxia in the tissue.
Streptococcus agalactiae (Group B Strep) requires zinc and nickel as critical cofactors and expresses beta-glucuronidase.
Fusobacterium nucleatum requires zinc and nickel as critical cofactors. It decreases local oxygen levels primarily by metabolizing and consuming oxygen. However, it is not directly involved in estrogen deconjugation.
Candida albicans deserves particular attention. I have heard researchers describe Candida in the context of endometriosis as an incidental finding and dismiss it as noise. Let me explain why that is wrong.
Candida biofilms consume oxygen, creating anaerobic pockets that allow obligate anaerobes like B. fragilis and F. nucleatum to persist. In a hypoxic state, facultative anaerobes switch to fermentative metabolism. There is solid evidence of tight cohesion and reduced cytokine output when Candida co-aggregates with F. nucleatum, suggesting functional shielding. Candida enhances epithelial colonization of Streptococcus. It creates mixed biofilms with E. coli and reduces host defenses. And nickel significantly increases the biomass of the biofilms in these co-aggregates.
This is not an incidental finding and this is not noise. This is a problem that 16S studies are not detecting. Researchers need to take microbiome signature findings a step further and look at the relationships between the taxa before dismissing anything as noise.
Lactobacillus presents an interesting pattern: enriched in the gut and peritoneum but decreased in the vaginal cavity. Translocation is likely the culprit, as the gut microbiome is a better predictor of endometriosis than the vaginal microbiome. Lactobacillus is also nickel-dependent and glyoxalase-positive.
Lachnospiraceae and Ruminococcaceae are likely decreased simply because they did not have the right defense systems to survive in the iron-rich environment.
Step 7: Dietary interventions through the framework lens
Clinically evaluated dietary interventions for endometriosis include a low red meat diet, a low-fat diet, a diet with increased citrus fruits, avoidance of brassica vegetables, and a low-nickel diet. Through the framework, each of these makes mechanistic sense.
A low red meat diet reduces free iron and zinc. A low-fat diet works well if E. coli is the major pathogen. Citrus fruits reduce circulating LPS and were found protective especially in smokers. Increased brassica consumption was found as a risk factor for endometriosis, and members of that family are cadmium, lead, zinc, and nickel hyperaccumulators, which are already elevated in the metallomic signature.
The low-nickel diet is particularly significant. Several studies have found that a low-nickel diet shows promise for endometriosis. But even the researchers involved in those studies have said they do not quite understand the mechanisms behind it.
The mechanism is glyoxalase. Glyoxalase positivity enables pathogen persistence because it enables them to evade neutrophils. Without glyoxalase, they become susceptible to immune responses. But glyoxalase is a nickel-dependent enzyme. Remove the nickel and you disarm the pathogens.
Step 8: Beneficial microbes as ecological interventions
Escherichia coli Nissle 1917 is one of my favorite probiotics. It lacks virulence genes. It outcompetes pathogenic strains of E. coli, B. fragilis, Group B Strep, F. nucleatum, Candida albicans, and other important pathogens including Klebsiella pneumoniae and Salmonella typhimurium. It has more sensitive siderophores and thus better iron acquisition systems. It also uptakes nickel and competes even in hypoxic environments.
Saccharomyces boulardii is a non-pathogenic fungal species that can outcompete Candida. Its cell walls can also bind cadmium and lead.
Step 9: Novel interventions predicted by the framework
Using this microbiome signature framework, one could easily have predicted that hyperbaric oxygen therapy (HBOT) would be beneficial due to the sheer number of strict and facultative anaerobes in the signature. HBOT was the only intervention I had come across in two years of looking that mentioned complete remission in an animal model. It has not been clinically investigated in humans yet. It probably should be.
Step 10: The STOP analysis
This is where the method produces its most clinically urgent output.
The following is established in the literature: increased dietary zinc and iron are associated with an increased risk of endometriosis. Lactoferrin and calprotectin, which sequester iron and zinc away from pathogens, are already elevated. Hepcidin is increased in endometriosis, which is an indicator of functional anemia, not true iron deficiency. Key pathogenic taxa express zinc-dependent metalloproteinases and advanced iron uptake systems that enable them to degrade host tissues and inhibit the host immune system.
Doctors often find that endometriosis patients have anemia or low serum zinc and prescribe iron and zinc supplementation. The clinical literature shows this exacerbates symptoms.
Does the routine recommendation of iron and zinc supplementation make sense based on this microbiome signature? No. This is a STOP: a Suggested Termination Of Practice. The host is deliberately restricting pathogen access to these metals through nutritional immunity. Supplementation overrides that defense and feeds the very organisms driving the disease.
The method, formalized
What I have just demonstrated is the Microbiome Signature Triangulation Method. The reasoning loop is:
Signature leads to mechanism, mechanism reveals vulnerability, vulnerability suggests intervention, intervention is validated against external clinical evidence, and any contradiction forces model revision.
An intervention is validated within this framework when three criteria are met simultaneously. First, there is credible evidence that the intervention moves at least one load-bearing signature feature in the predicted direction (target validation). Second, there is credible evidence that the intervention improves relevant clinical endpoints in the target condition (clinical validation). Third, the intervention's effects do not violate the ecological and physiological story induced by the full signature (system coherence).
When all three criteria are met, the intervention is validated and the underlying microbiome signature is simultaneously confirmed as clinically actionable. When any criterion fails, the model is wrong and must be revised.
This is a self-falsifying system. That is by design. It means the method gets stronger with every contradiction it encounters, because contradictions force revisions that make the model more accurate.
What this means for the field
The conventional timeline for microbiome science reaching clinical application is estimated at 30 to 50 years. I have been doing this for 15.
The bottlenecks that the field cites (lack of high-quality RCTs, lack of standardization, complexity of the microbiome, insufficient mechanistic understanding) are real, but they are not obstacles to translational work. They are obstacles to a specific kind of translational work: the kind that requires all variance to be eliminated before anyone acts.
The method described here works differently. It tolerates regional variance. It incorporates inconsistent methods. It uses mechanism and triangulation to generate actionable hypotheses from imperfect data. And it produces outputs that can be tested: specific dietary interventions, specific probiotics, specific novel therapies, and specific practices that should be stopped.
A non-exhaustive microbiome signature was enough to shape a clinical recommendation in 15 minutes at a conference. That same reasoning system, applied systematically across conditions, is what the Microbiome Signatures Database is built to scale.